Regulation and Control of Metabolism in Bacteria (page 3)
(This chapter has 5 pages)
© Kenneth Todar, PhD
Enzyme Repression
Enzyme repression is a form of negative control
(down-regulation)
of bacterial transcription. This process, along with that of enzyme
induction,
is called negative control because a regulatory protein brings
about
inhibition
of mRNA synthesis which leads to decreased synthesis of
enzymes.
Although feedback inhibition shuts off synthesis of the end product
of a pathway, it still allows some waste of energy and carbon if the
cell
continued to manufacture enzymes for which it has no use. It is the
process
of enzyme repression that prevents the synthesis of the enzymes
concerned
with the synthesis of that particular end product. In the case of
the
pathway of tryptophan biosynthesis (Figure 3), the end product of the
pathway,
tryptophan, serves as an effector molecule that can shutdown the
synthesis
of the Enzymes a, b, c,
d, and e that are concerned with tryptophan
biosynthesis.
This results in saving of many molecules of ATP which must be expended
during protein synthesis, and it conserves amino acid precursors for
synthesis
of other proteins. The process is slower to act than is feedback
inhibition
(which acts immediately) because pre-existing enzymes have to be
diluted
out as a result of cell division before its effects are seen.
The genes for tryptophan biosynthesis in Escherichia coli are
organized on the bacterial chromosome in the tryptophan operon (trp
operon). An operon is a
cluster of genes that are controlled by the
same elements and which are coordinately transcribed and translated.
The
trp operon consists of a Promoter (P) region, an Operator (O) region,
an
Attenuator (A) region, and the five structural genes for the enzymes
involved
in
tryptophan biosynthesis (Trp A-E) The components of the trp operon and
its control elements are described in Figure 5 and Table 2 below.
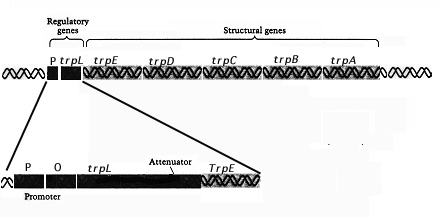
Figure 5. Genetic organization
of the Trp operon and its control elements.
Table 2. The Trp operon and
its control elements
R = Regulatory gene that
encodes
for the trp Repressor protein that is concerned with regulating the
synthesis
of the 5 gene products. An active repressor binds to a specific
nucleotide
sequence in the operator region and thereby blocks binding of RNAp to
the
promoter to initiate transcription.
O = Operator specific
nucleotide
sequence on DNA to which an active Repressor binds.
P = Promoter specific
nucleotide
sequence on DNA to which RNA polymerase binds to initiate
transcription.
If the repressor protein binds to the operator, RNAp is prevented from
binding with the promoter and initiating transcription. Therefore, none
of the enzymes concerned with tryptophan biosynthesis are synthesized.
A = Attenuator DNA sequence
which
lies between the operator and the structural genes for trp
biosynthesis.
The attenuator is a barrier that RNA polymerase must traverse if it is
to transcribe the genes for tryptophan biosynthesis. In the presence of
trp, most RNAp molecules fall off the DNA before transcribing the trp
genes.
In the absence of trp, RNAp is able to traverse the attenuator region
to
successfully transcribe the trp genes.
Trp A, B, C, D, E =
Structural genes for enzymes involved in tryptophan biosynthesis.
Trp = tryptophan end
product
of the biosynthetic pathway. When combined with the repressor
protein the Repressor is active. Trp is called a corepressor.
The trp operon is regulated by a regulatory gene (Trp L) associated
with
the trp promoter. The product of the Trp L gene is the trp Repressor,
an allosteric protein which is regulated by tryptophan. The Repressor
is
produced constitutively in small amounts in an inactive form. When the
Repressor combines with tryptophan it becomes activated and binds to
the
DNA of the trp operon in such a way that it blocks the transcription of
the structural genes for tryptophan. Thus, in the presence of
tryptophan,
transcription of the genes for tryptophan biosynthesis are repressed
(tryptophan
is not produced), while in the absence of tryptophan, the genes for
tryptophan
biosynthesis can be transcribed (tryptophan is produced); See Figure 6
below.

Figure 6a. Derepression of the
trp operon. In the absence of trp the inactive repressor cannot bind to
the operator to block transcription. The cell must synthesize the amino
acid.

Figure 6b. Repression of the
trp operon. In the presence of tryptophan the trp operon is repressed
because
trp activates the repressor. Transcription is blocked because the
active
repressor binds to the DNA and prevents binding of RNA polymerase.
chapter continued
Previous Page